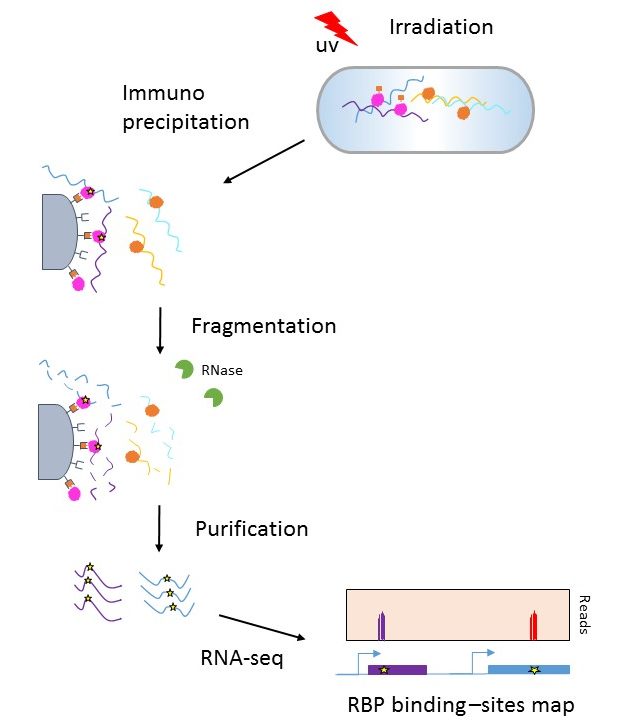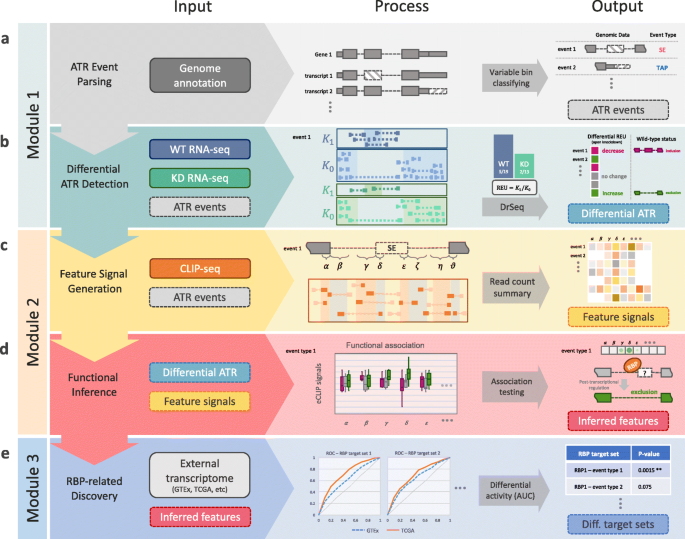
SURF: integrative analysis of a compendium of RNA-seq and CLIP-seq datasets highlights complex governing of alternative transcriptional regulation by RNA-binding proteins | Genome Biology | Full Text

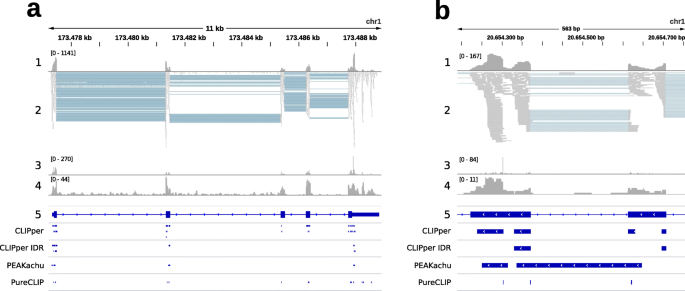
CLIP-seq data for different RBPs, CLIP technologies, and peak calling... | Download Scientific Diagram
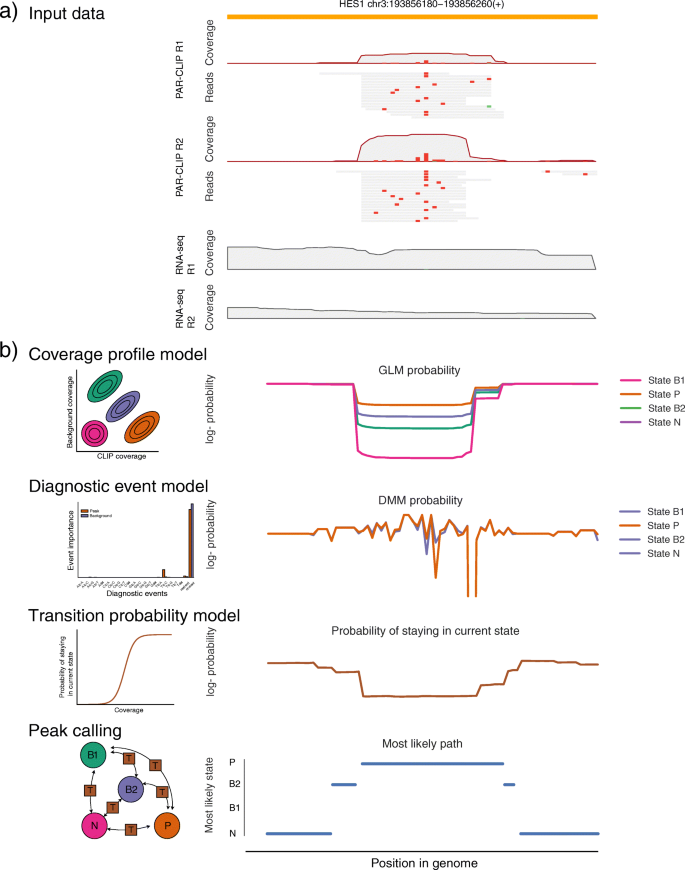
omniCLIP: probabilistic identification of protein-RNA interactions from CLIP -seq data | Genome Biology | Full Text

omniCLIP: probabilistic identification of protein-RNA interactions from CLIP -seq data | RNA-Seq Blog

Summary of the analysis software, pipelines and databases for CLIP-Seq... | Download Scientific Diagram
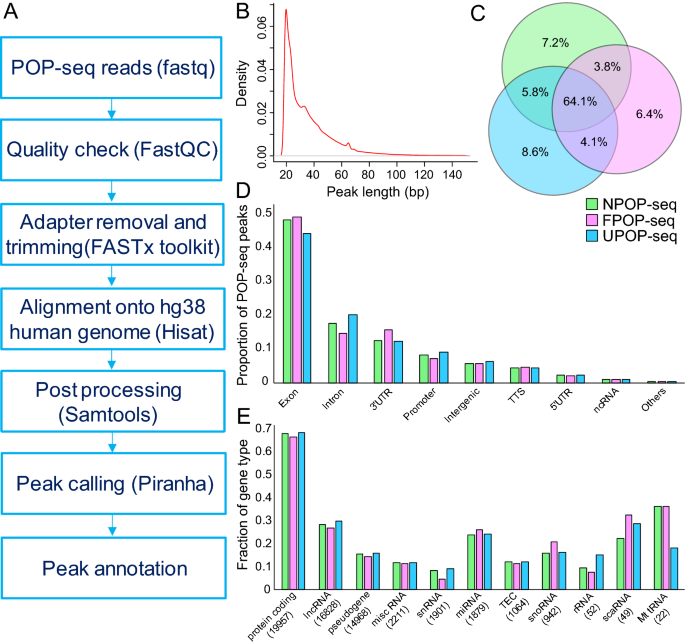
Transcriptome-wide high-throughput mapping of protein–RNA occupancy profiles using POP-seq | Scientific Reports

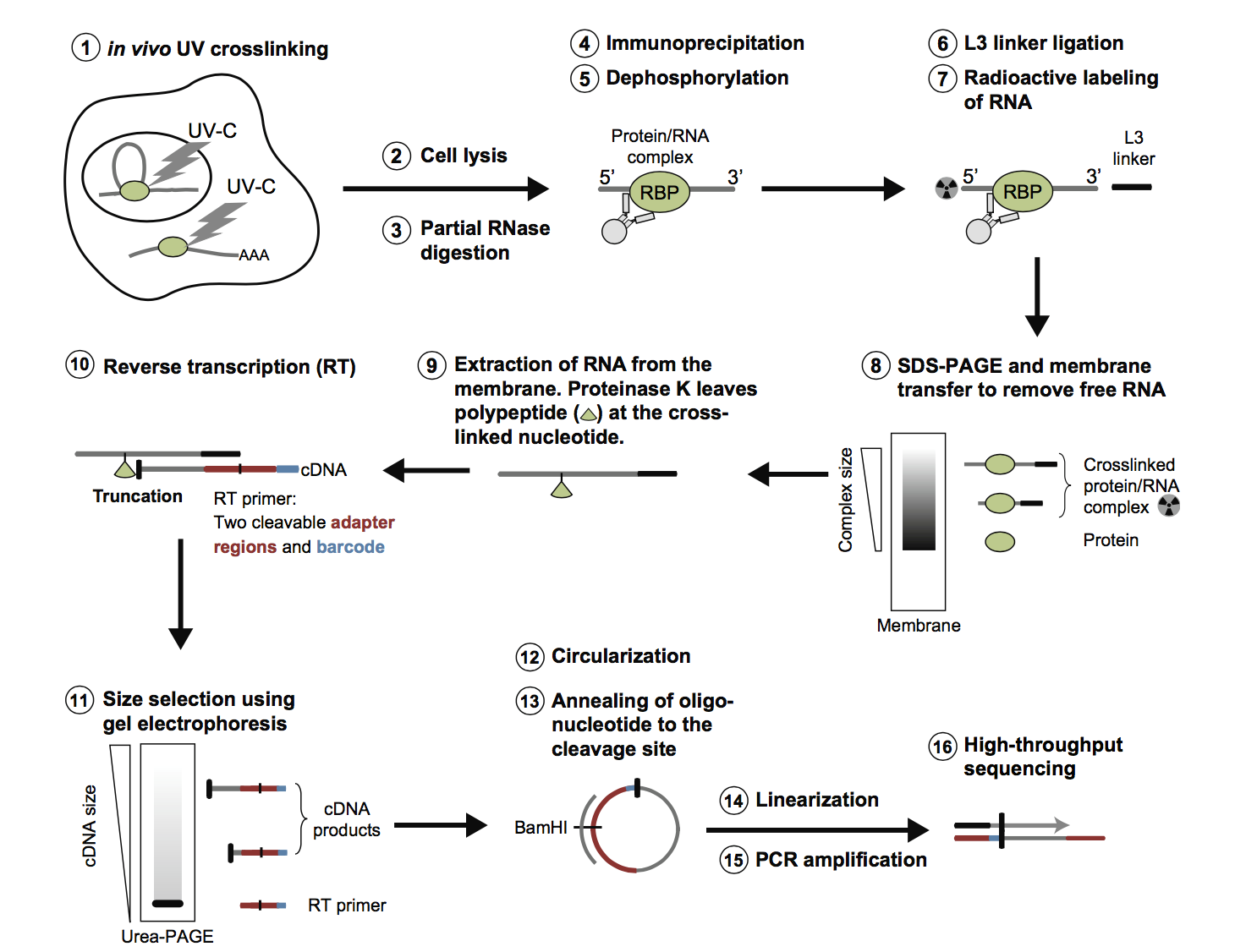


![PDF] Current methods in the analysis of CLIP-Seq data | Semantic Scholar PDF] Current methods in the analysis of CLIP-Seq data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e504d75c2c48b19613a796fb54083b71e3781289/2-Figure1-1.png)
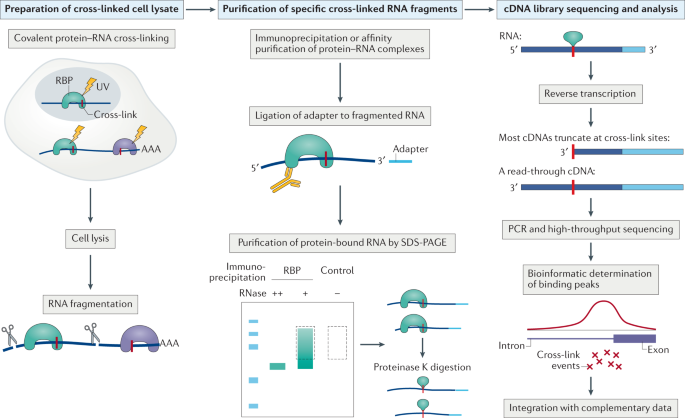


![PDF] CLIPSeqTools—a novel bioinformatics CLIP-seq analysis suite | Semantic Scholar PDF] CLIPSeqTools—a novel bioinformatics CLIP-seq analysis suite | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6bbd241680ec0f3d6a86dec738a6b864ea5048c3/6-Figure3-1.png)
![PDF] CLIPSeqTools—a novel bioinformatics CLIP-seq analysis suite | Semantic Scholar PDF] CLIPSeqTools—a novel bioinformatics CLIP-seq analysis suite | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6bbd241680ec0f3d6a86dec738a6b864ea5048c3/2-Figure1-1.png)


![PDF] Galaxy CLIP-Explorer: a web server for CLIP-Seq data analysis | Semantic Scholar PDF] Galaxy CLIP-Explorer: a web server for CLIP-Seq data analysis | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/450fd3c214476c2fa46ac3697732d0b5704f8eec/6-Figure1-1.png)