Learning to Predict miRNA-mRNA Interactions from AGO CLIP Sequencing and CLASH Data | PLOS Computational Biology

Computational approaches for the analysis of RNA–protein interactions: A primer for biologists - ScienceDirect

CLIP-Seq and massively parallel functional analysis of the CELF6 RNA binding protein reveals a role in destabilizing synaptic gene mRNAs through interaction with 3'UTR elements in vivo | bioRxiv

Pipeline for extracting MIWI CLIP-Seq features (see Section 2). The RNA... | Download Scientific Diagram

Summary of the analysis software, pipelines and databases for CLIP-Seq... | Download Scientific Diagram

circTools: identify and annotate circRNAs and their interactions with microRNAs (miRNAs) from large-scale <strong>CLIP-Seq (HITS-CLIP, PAR-CLIP, iCLIP, CLASH)</strong> and RNA-seq data.
PRAS: Predicting functional targets of RNA binding proteins based on CLIP- seq peaks | PLOS Computational Biology
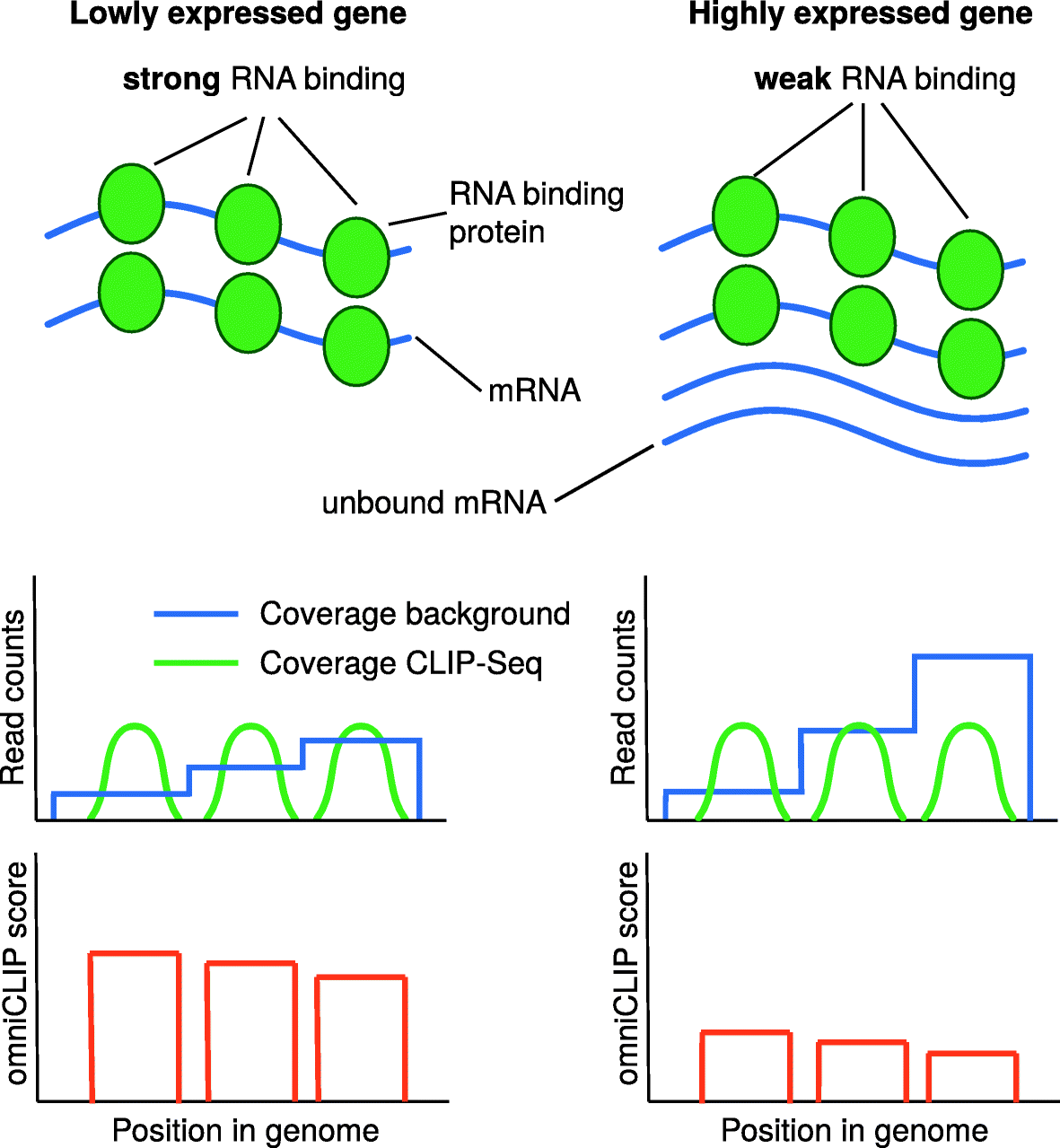
omniCLIP: probabilistic identification of protein-RNA interactions from CLIP -seq data | Genome Biology | Full Text


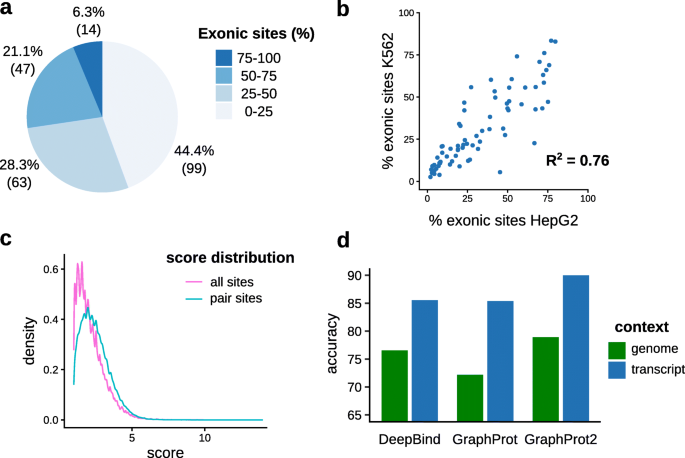
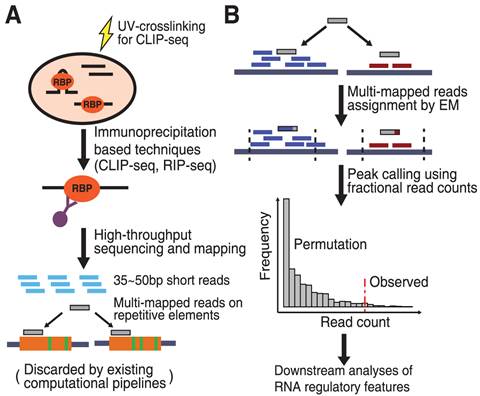
![PDF] CLIPSeqTools—a novel bioinformatics CLIP-seq analysis suite | Semantic Scholar PDF] CLIPSeqTools—a novel bioinformatics CLIP-seq analysis suite | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6bbd241680ec0f3d6a86dec738a6b864ea5048c3/2-Figure1-1.png)
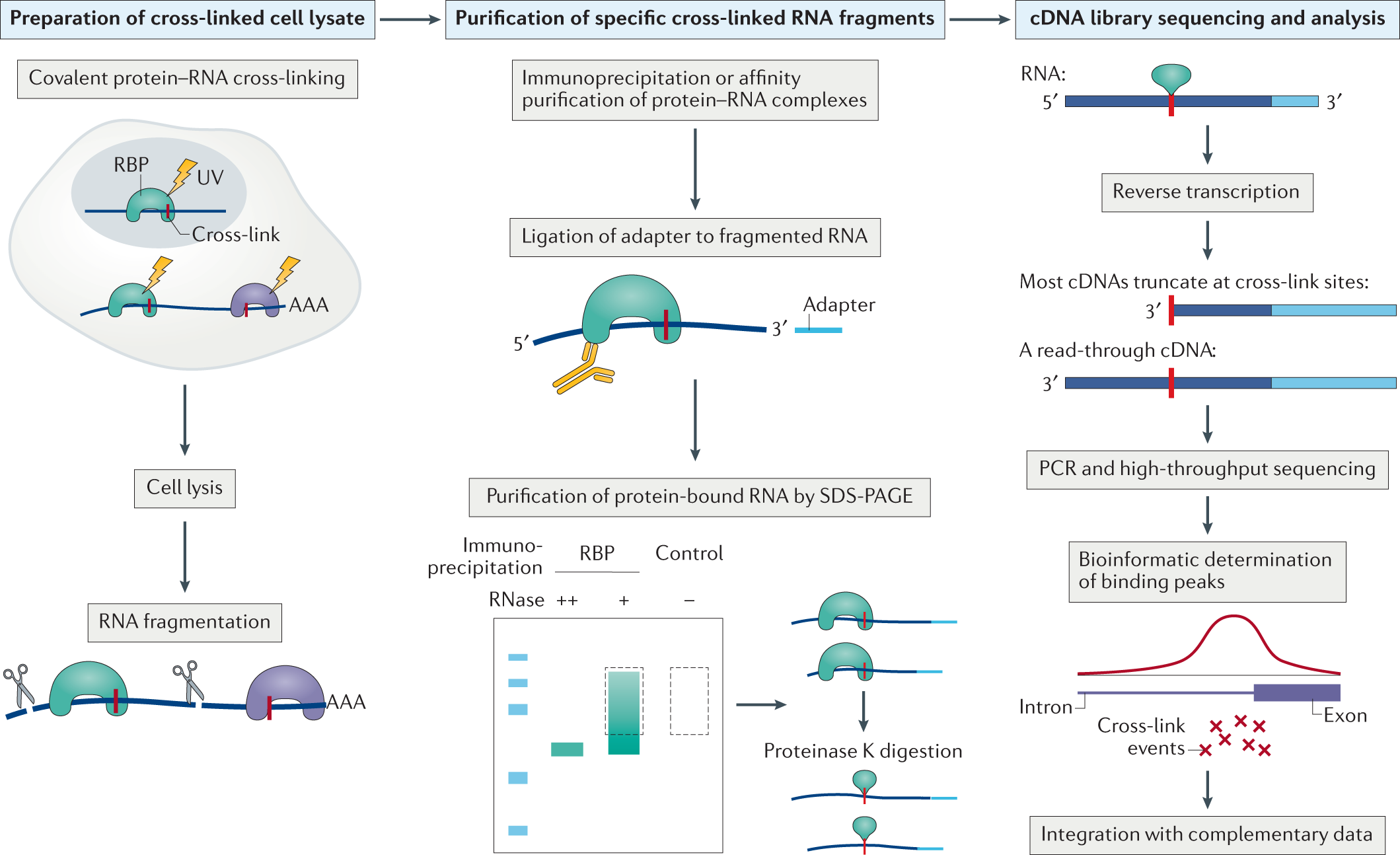

![PDF] CLIPSeqTools—a novel bioinformatics CLIP-seq analysis suite | Semantic Scholar PDF] CLIPSeqTools—a novel bioinformatics CLIP-seq analysis suite | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6bbd241680ec0f3d6a86dec738a6b864ea5048c3/6-Figure3-1.png)
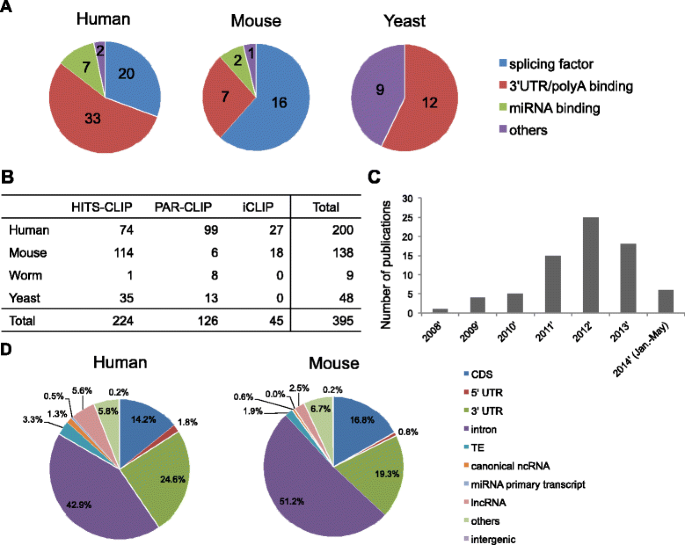
![PDF] Current methods in the analysis of CLIP-Seq data | Semantic Scholar PDF] Current methods in the analysis of CLIP-Seq data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e504d75c2c48b19613a796fb54083b71e3781289/3-Figure2-1.png)
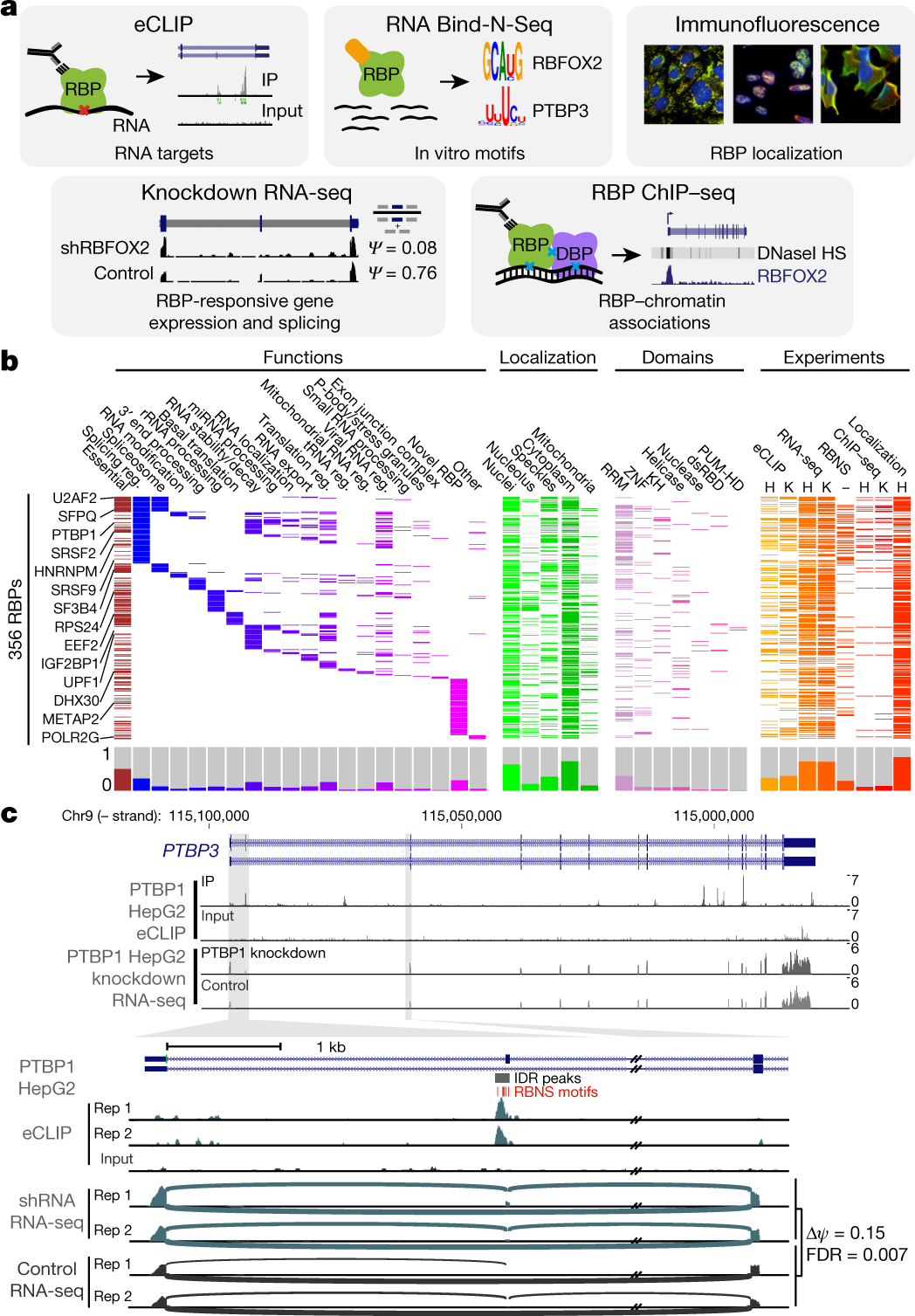
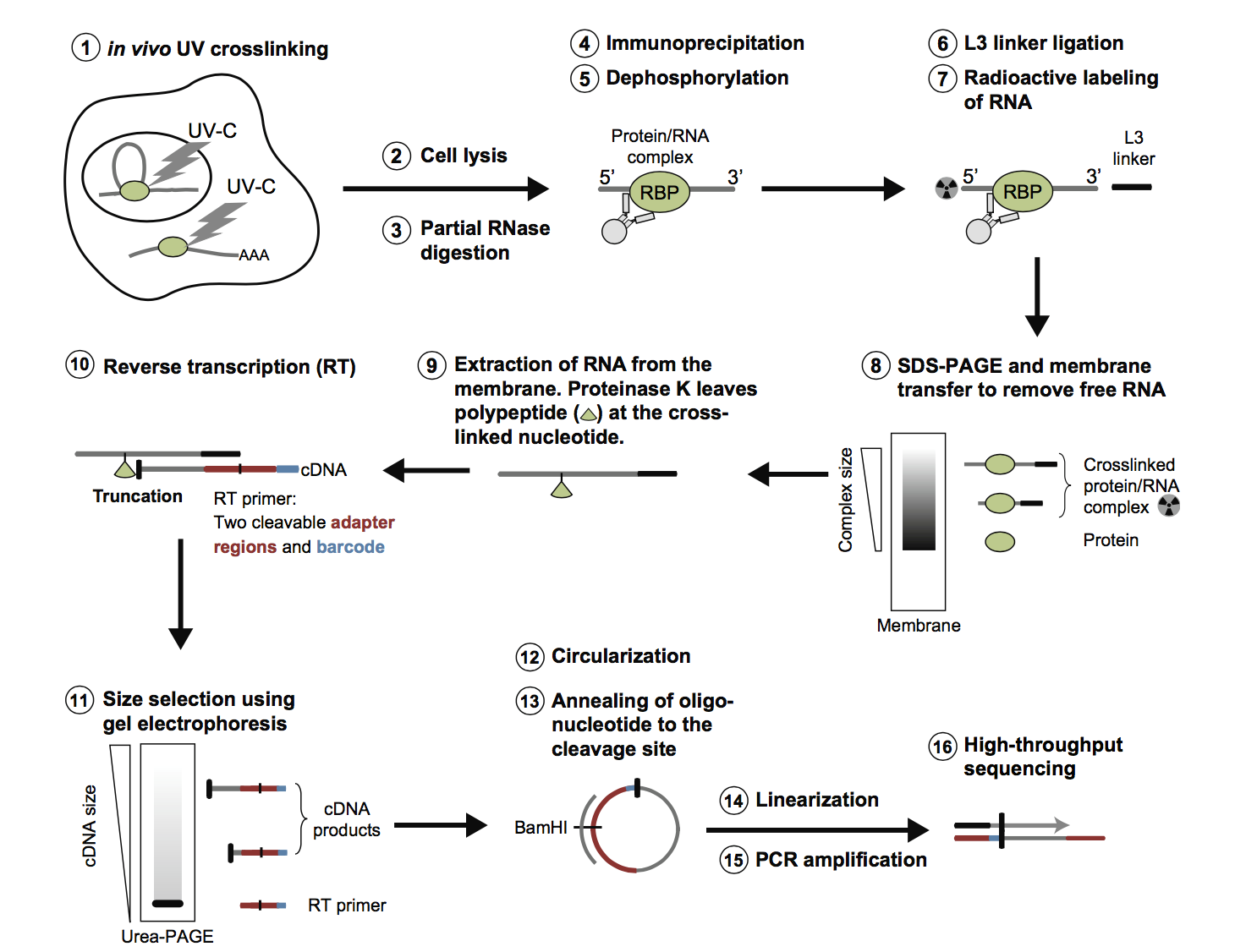




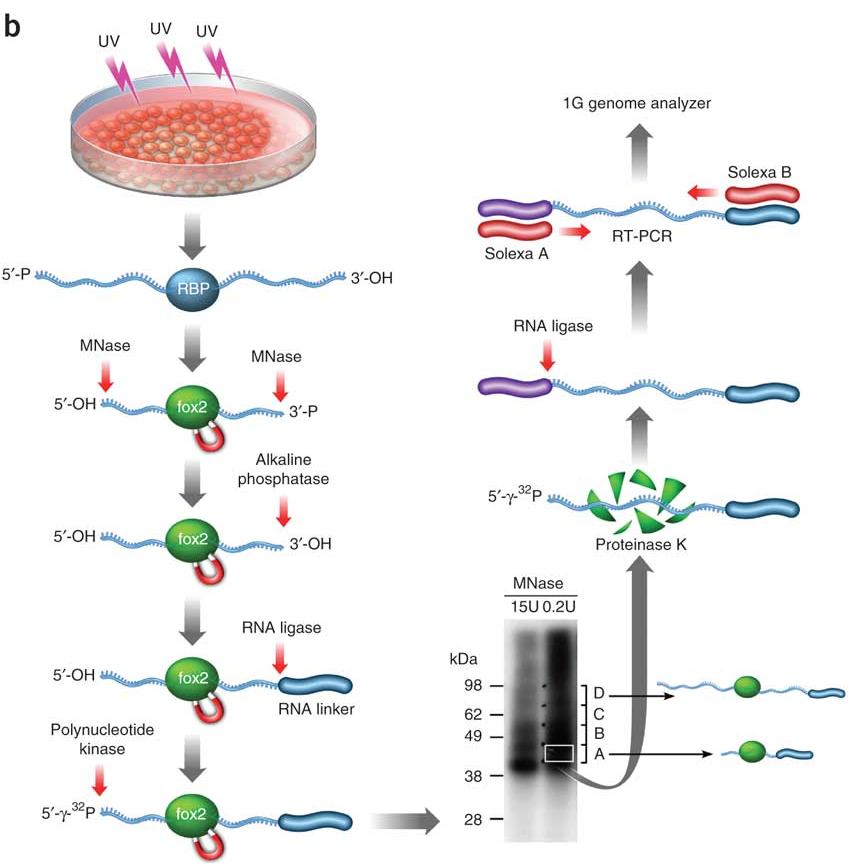

![PDF] Current methods in the analysis of CLIP-Seq data | Semantic Scholar PDF] Current methods in the analysis of CLIP-Seq data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e504d75c2c48b19613a796fb54083b71e3781289/2-Figure1-1.png)